DNA: Double Helix
- Page ID
- 467
The secondary structure of DNA is actually very similar to the secondary structure of proteins. The protein single alpha helix structure held together by hydrogen bonds was discovered with the aid of X-ray diffraction studies. The X-ray diffraction patterns for DNA show somewhat similar patterns.
Introduction
In addition, chemical studies by E. Chargaff indicate several important clues about the structure of DNA. In the DNA of all organisms:
- Chargaff's findings clearly indicate that some type of heterocyclic amine base pairing exists in the DNA structure. X-ray diffraction data shows that a repeating helical pattern occurs every 34 Angstrom units with 10 subunits per turn. Each subunit occupies 3.4 Angstrom units which is the same amount of space occupied by a single nucleotide unit. Using Chargaff's information and the X-ray data in conjunction with building actual molecular models, Watson and Crick developed the double helix as a model for DNA.
The double helix in DNA consists of two right-handed polynucleotide chains that are coiled about the same axis. The heterocyclic amine bases project inward toward the center so that the base of one strand interacts or pairs with a base of the other strand. According to the chemical and X-ray data and model building exercises, only specific heterocyclic amine bases may be paired.
Base Pairing Principle
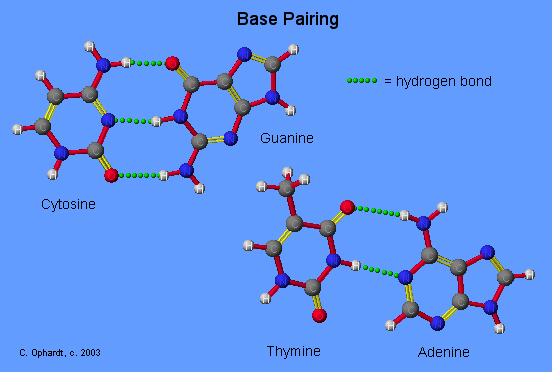
The Base Pairing Principle is that adenine pairs with thymine (A - T) and guanine pairs with cytosine (G - C)
The base pairing is called complementary because there are specific geometry requirements in the formation of hydrogen bonds between the heterocylic amines. Heterocyclic amine base pairing is an application of the hydrogen bonding principle. In the structures for the complementary base pairs given in the graphic on the left, notice that the thymine - adenine pair interacts through two hydrogen bonds represented as (T=A) and that the cytosine-guanine pair interacts through three hydrogen bonds represented as (C=G).
Although other base pairing-hydrogen bonding combinations may be possible, they are not utilized because the bond distances do not correspond to those given by the base pairs already cited. The diameter of the helix is 20 Angstroms.
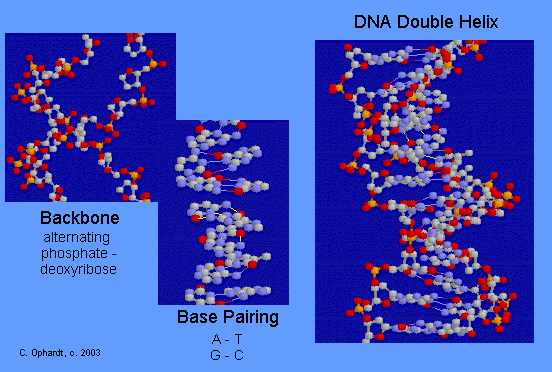
DNA Double Helix
The double-stranded helical model for DNA is shown in the graphic on the left. The easiest way to visualize DNA is as an immensely long rope ladder, twisted into a cork-screw shape. The sides of the ladder are alternating sequences of deoxyribose and phosphate (backbone) while the rungs of the ladder (bases) are made in two parts with each part firmly attached to the side of the ladder. The parts in the rung are heterocyclic amines held in position by hydrogen bonding. Although most DNA exists as open ended double helices, some bacterial DNA has been found as a cyclic helix. Occasionally, DNA has also been found as a single strand.
Problems
QUES. Describe the structure of the double helix of DNA in your own words including the terms: backbone, heterocyclic amines, complementary base pairings, hydrogen bonding, deoxyribose, phosphate.
Quiz: In RNA, which base hydrogen bonds with uracil? Carefully compare the structure of uracil to the others to find the one that is most similar. Quiz: If DNA is heated, what happens to the double helix? Hint: The result is similar to the denaturing of a protein by the same method. What type of bonding holds the secondary structure of both proteins and DNA? Outside Links
- The structure of the 'Dickerson Dodecamer' was originally reported in: Drew, H. R., Wing, R. M., Takano, T., Broka, C., Tanaka, S., Itakura, K. & Dickerson, R. E. (1981). Structure of a B-DNA dodecamer: conformation and dynamics. Proc. Natl. Acad. Sci. USA 78, 2179-2183. and the coordinates were acquired from the Brookhaven Protein Data Bank. The filename is 1BNA.
Contributors
- Charles Ophardt, Professor Emeritus, Elmhurst College; Virtual Chembook

