9.1: Methyl group transfers: examples of SN2 reactions
- Page ID
- 1090
9.1A: SAM methyltransferases
Some of the most important examples of SN2 reactions in biochemistry are those catalyzed by S-adenosyl methionine (SAM) – dependent methyltransferase enzymes. We have already seen, in chapter 6 and again in chapter 8, how a methyl group is transferred in an SN2 reaction from SAM to the amine group on the nucleotide base adenosine:
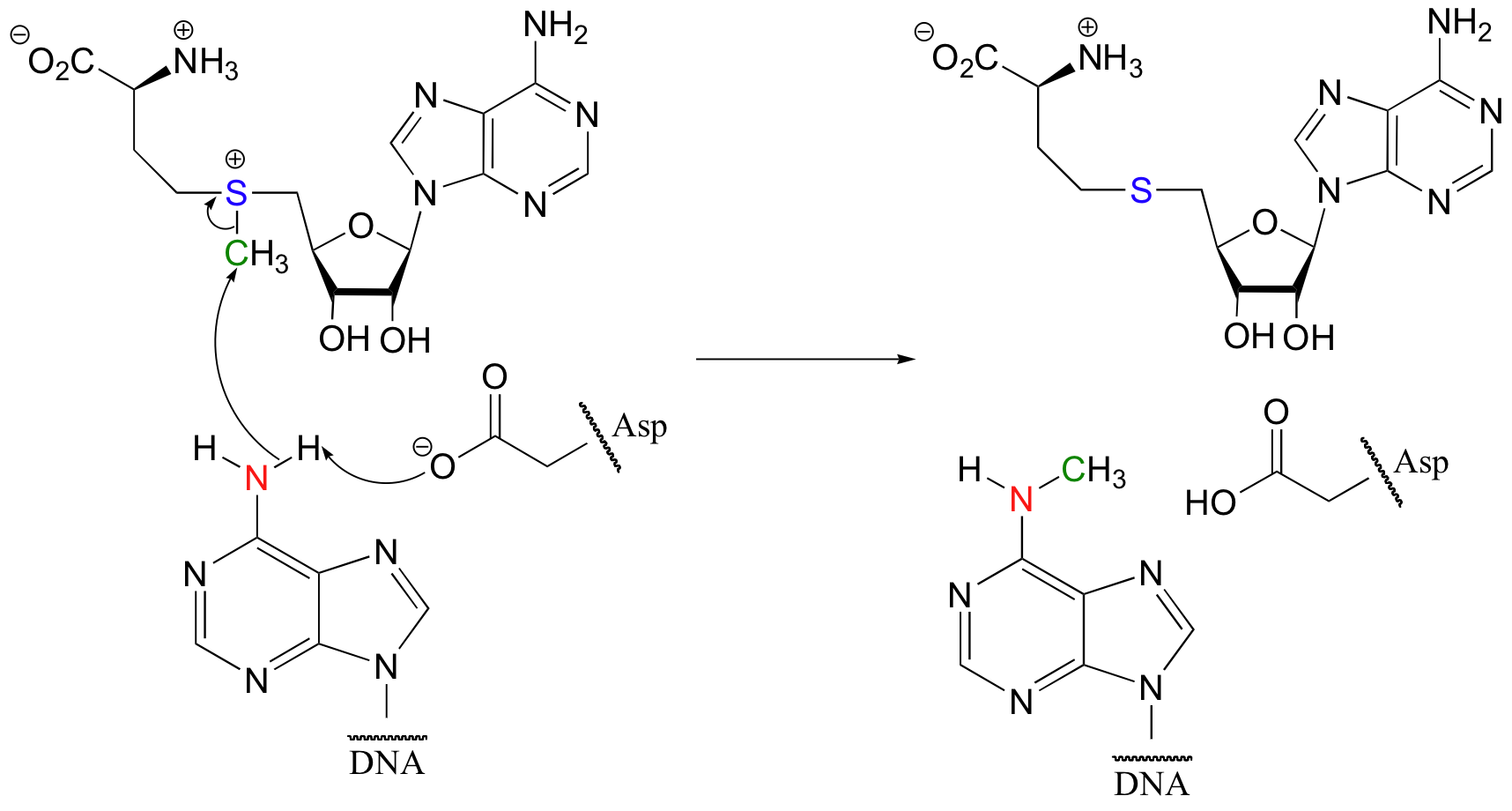
(Nucleic Acids Res. 2000, 28, 3950).
Another SAM-dependent methylation reaction is catalyzed by an enzyme called catechol-O-methyltransferase. The substrate here is epinephrine, also known as adrenaline.
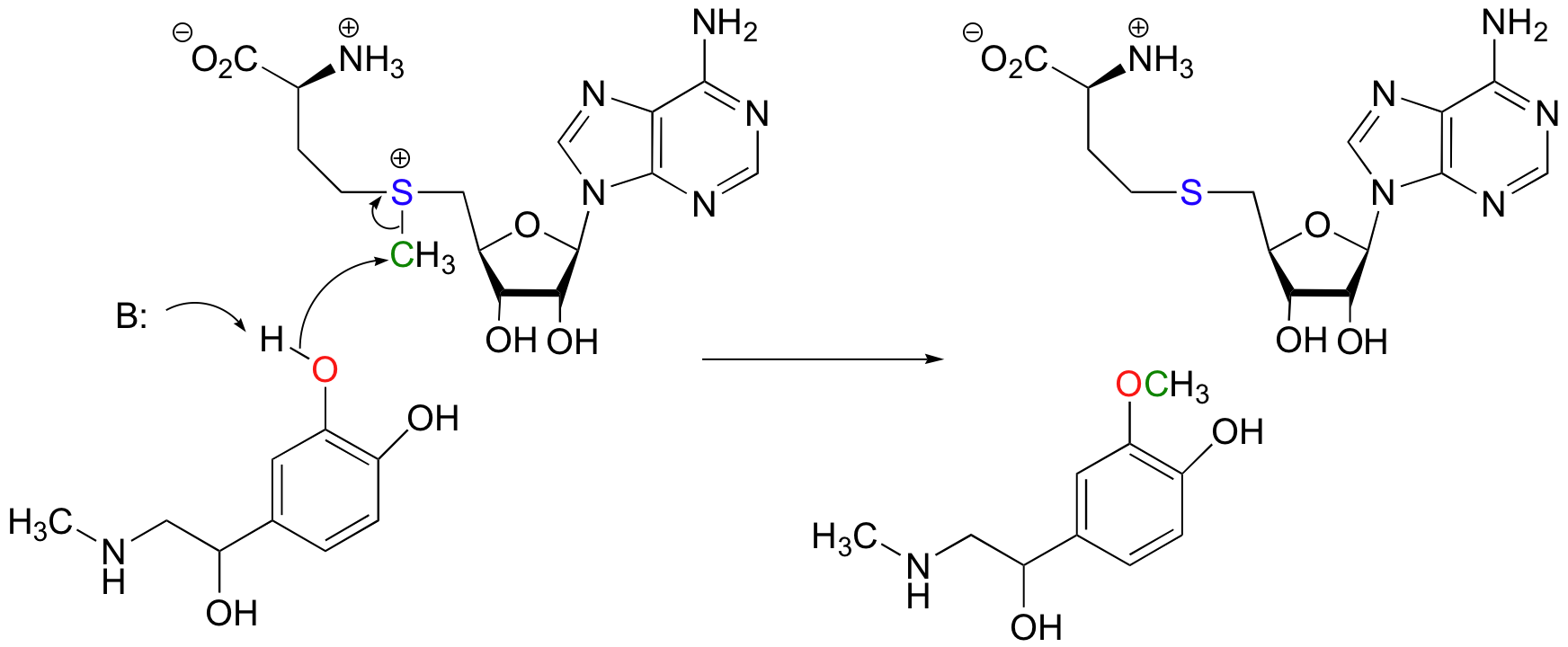
Notice that in this example, the attacking nucleophile is an alcohol rather than an amine (that’s why the enzyme is called an O-methyltransferase). In both cases, though, a basic amino acid side chain is positioned in the active site in just the right place to deprotonate the nucleophilic group as it attacks, increasing its nucleophilicity. The electrophile in both reactions is a methyl carbon, so there is little steric hindrance to slow down the nucleophilic attack. The methyl carbon is electrophilic because it is bonded to a positively-charged sulfur, which is a powerful electron withdrawing group. The positive charge on the sulfur also makes it an excellent leaving group, as the resulting product will be a neutral and very stable sulfide. All in all, in both reactions we have a reasonably good nucleophile, an electron-poor, unhindered electrophile, and an excellent leaving group.
Because the electrophilic carbon in these reactions is a methyl carbon, a stepwise SN1-like mechanism is extremely unlikely: a methyl carbocation is very high in energy and thus is not a reasonable intermediate to propose. We can confidently predict that this reaction is SN2. Does this SN2 reaction occur, as expected, with inversion of stereochemistry? Of course, the electrophilic methyl carbon in these reactions is achiral, so inversion is not apparent. To demonstrate inversion, the following experiment has been carried out with catechol-O-methyltransferase:
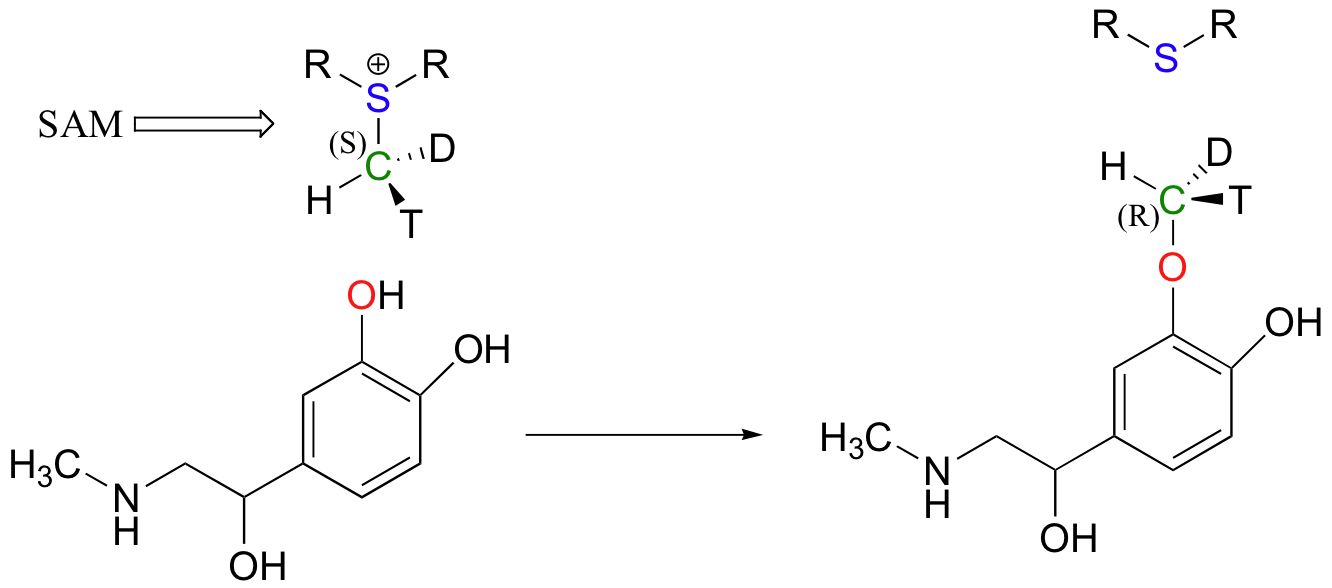
Here, the methyl group of SAM was made to be chiral by incorporating hydrogen isotopes tritium (3H, T) and deuterium (2H, D). The researchers determined that the reaction occurred with inversion of configuration, as expected for an SN2 displacement (J. Biol. Chem. 1980, 255, 9124).
Exercise 9.1: SAM is formed by a nucleophilic substitution reaction between methionine and adenosine triphosphate (ATP). Propose a mechanism for this reaction.
Callstack:
at (Under_Construction/Purgatory/Book:_Organic_Chemistry_with_a_Biological_Emphasis_(Soderberg)/Chapter_09:_Nucleophilic_substitution_reactions_II/9.1:_Methyl_group_transfers:_examples_of_SN2_reactions), /content/body/div[1]/div[3]/span, line 1, column 20
9.1B: Synthetic parallel – the Williamson ether synthesis
Synthetic organic chemists often use reactions in the laboratory that are conceptually very similar to the SAM-dependent methylation reactions described above. In what is referred to as "O-methylation", an alcohol is first deprotonated by a strong base, typically sodium hydride (this is essentially an irreversible deprotonation, as the hydrogen gas produced can easily be removed from the reaction vessel).
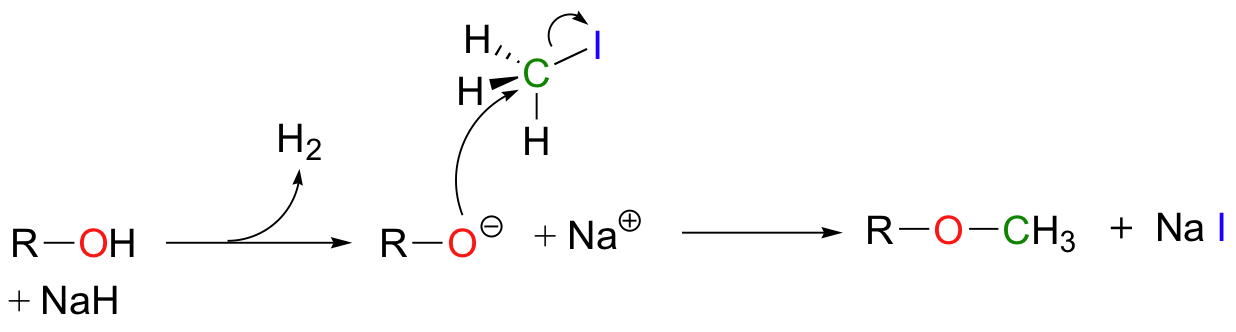
The alkoxide ion, a powerful nucleophile, is then allowed to attack the electrophilic carbon in iodomethane, displacing iodide in an SN2 reaction. Recall (section 7.3A) that iodide ion is the least basic - and thus the best leaving group - among the halogens commonly used in the synthetic lab. CH3I is one of the most commonly used lab reagents for methyl transfer reactions, and is the lab equivalent of SAM. In the synthesis of a modified analog of the signaling molecule myo-inisitol triphosphate, an alcohol group was methylated using sodium hydride and iodomethane (Carbohydrate Research 2004, 339, 51):
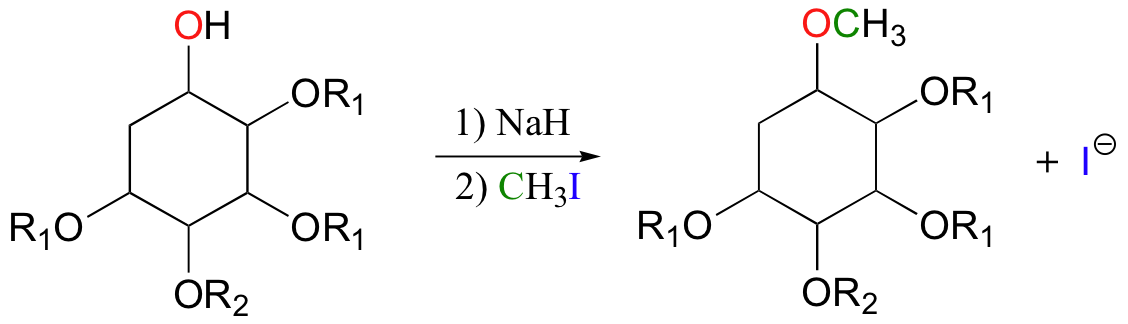
Methylation of amines (N-methylation) by iodomethane is also possible. A recent study concerning the optimization of peptide synthesis methods involved the following reaction (J. Org. Chem. 2000 65, 2309):
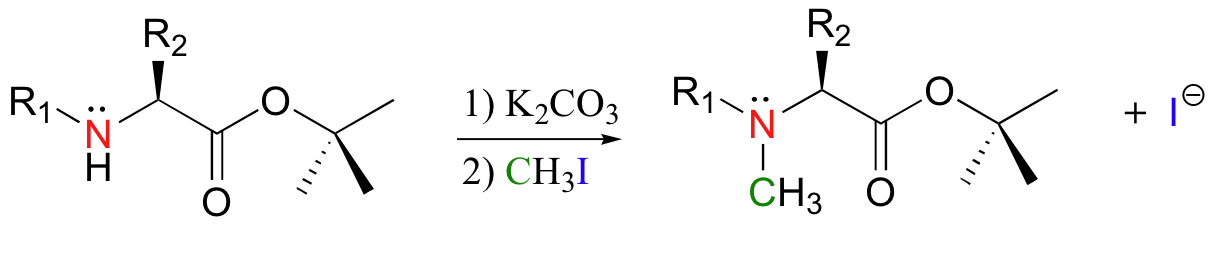
Exercise 9.2: Notice that in the N-methylation reaction a weak base (potassium carbonate) is used, rather than the very strong base (sodium hydride) that was used in the o-methylation reaction. Why is this weak base sufficient for N-methylation but not for o-methylation? In your answer, draw a step-by step mechanism for both reactions, and think carefully about pKa values and where in each mechanism the deprotonation step occurs.
Solution
The methylation of an alcohol by iodomethane is just one example of what is generally referred to as the Williamson ether synthesis, in which alcohols are first deprotonated by a strong base and then allowed to attack an alkyl halide electrophile. Below we see the condensation between ethanol and benzyl bromide:
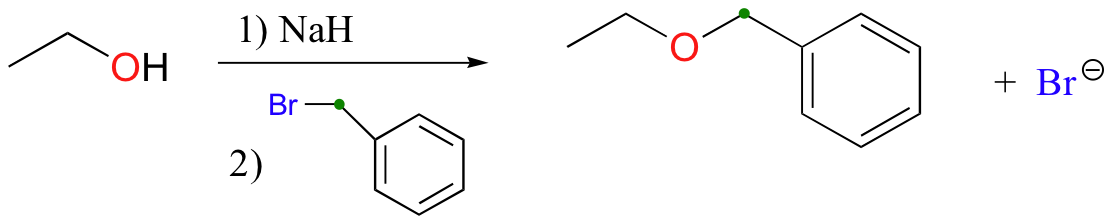
It is important to note that this reaction does not work well at all if secondary or tertiary alkyl halides are used: remember that the alkoxide ion is a strong base as well as a nucleophile, and elimination will compete with nucleophilic substitution (Section 8.5D).


